Heriot-Watt Complex Systems Laboratory
Controlling Genetic Networks
Genetic regulatory networks (GRNs) are responsible for the behaviour of biological cells. Diseases such as cancer occur when GRNs enter abnormal parts of their state space. We are looking at how synthetic GRNs can be used to control the behaviour of a native biological GRN, guiding it to particular parts of its state space. Currently this work is being done in simulation, but eventually our aim is to develop synthetic biology implementations within real cells.
Artificial Biochemical Networks

Biochemical networks underlie the complexity of biological organisms. We have been looking at the properties of connectionist architectures whose structure and function are motivated by the organisation of biochemical networks. We have found these architectures to be an effective means of solving difficult signal processing and control problems, particularly when interacting with real world complex dynamical systems.
Predictive Models of Parkinson's Disease

Parkinson's disease is a chronic neurodegenerative condition with a high incidence. We are applying advanced bio-inspired machine learning methods in order to build models of the underlying movement disorders associated with the disease, and using these models to assist clinicians when diagnosing and monitoring patients. This work is being done in collaboration with a number of leading medical centres around the world.
Bio-Inspired Data Mining
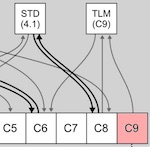
Biological systems have various characteristics that make them well suited to identifying patterns in information, and many of these characteristics are captured within bio-inspired computing models. Focusing on the use of evolutionary algorithms, cellular automata, and connectionist architectures, we have demonstrated how bio-inspired methods may be applied to various problems in data mining and knowledge discovery.