Below list summarises ongoing and recent research projects. Further details can be found on my research group's, i.e. the BISEL, web site. The corresponding BISEL links are listed on the left.
PhenoImageShare2013 - 2016, BBSRC-funded, Principal Investigator
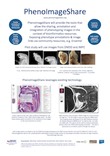
Summary: In collaboration with the MRC HGU and EBI, BISEL have embarked upon the BBSRC funded PhenoImageShare project. PhenoImageShare is creating a prototype infrastructure to support the sharing of phenotype annotations on phenotype images.
Further details: [PhenoImageShare Web Site]
Atlas Informatics
since 2012, Visiting Biomedical Informatics Research Scientist;
2001 - 2012, Biomedical Informatics Research Scientist, p/t secondment, MRC-funded;Summary: The Mouse Atlas project at MRC HGU provides a spatio-temporal framework for genomic and proteomic data about the developing mouse embryo. It consists of EMAP, the actual 3D mouse atlas and anatomy ontology, and EMAGE, a database of in-situ gene expression data mapped onto the atlas. As part of my work at the MRC I am involved in research and development of interoperability solutions for the Mouse Atlas.
Further Details: [Mouse Atlas Project Web Site]
CUBIST: Combining and Uniting Business Intelligence with Semantic Technologies
2010 - 2013, ICT EU-funded, Principal Investigator;Summary: The goal of CUBIST is to develop new Business Intelligence (BI) solutions, specifically through the application of Semantic Web technologies, to address the challenge of every increasing amounts of non-structured data; traditionally, BI only deals with structured data. At Heriot-Watt, we focus on a biomedical use case for CUBIST, in particular the semantic representation of image data in the Mouse Atlas and its exploitation through the CUBIST system.
Further Details: [CUBIST on BISEL pages] [CUBIST project web site]
RICORDO: Interoperable Anatomy and Physiology
2010 - 2012, ICT EU-funded; Principal Investigator;Summary: RICORDO was part of the EU's Virtual Physiological Human (VPH) programme and worked on a communal ontology-based annotation and repository communication strategy that supports the interoperability of VPH data and models across different biological scales. I was co-leader of the work package on interoperability of volumetric data, that dealt with integration of biomedical atlases and other 3D and 4D data and models in VPH.
Further Details: [RICORDO on BISEL pages] [RICORDO project web site]
INCF: Digital Atlasing Program
since 2008, INCF-funded; Task Force Member and Principal Investigator for Pilot Study (4/09-10/09);Summary: Digital atlases have been identified as potential neuroinformatics frameworks, as they act as a two-dimensional or three-dimensional map that one may use to traverse the brain and associated data of different granularity, modalities, and sources. The Task Force investigates the state of the art of rodent brain digital atlasing and formulates standards, guidelines, and policy recommendations.
A spin-off from this activity was a pilot project investigating the integration between the Edinburgh Mouse Atlas Project (EMAP) and the Allen Brain Atlas (ABA). Specifically, in BISEL we explored the use of NIF integration infrastructure. More recently we were involved in the development of a INCF hub for the Edinburgh Mouse Atlas Project as part of the INCF Distributed Atlas Infrastructure (INCF-DAI).
Further Details: [INCF on BISEL pages] [INCF web site]
Argudas: An Argumentation System for Gene Expression Databases
2009 - 2011, BBSRC-funded; Principal Investigator;Summary: The objective of the Argudas project was the development and deployment of a web-based argumentation tool that allows biologists to discover and assess inconsistencies and incompleteness within and across multiple gene expression databases. It presents the user with arguments for and against specific gene expression statements and, if requested by the user, will weigh up the evidence from the different databases to suggest an overall conclusion with respect to whether or not a particular gene is expressed in a given anatomical structure.
Further Details: [Argudas on BISEL pages]
Sealife: A Semantic Grid Browser for the Life Sciences
2006 - 2009, ICT EU-funded; Principal Investigator;Summary: The objective of Sealife was the conception and realisation of a semantic Grid browser for the Life Sciences, which links the existing Web to the currently emerging eScience infrastructure. At Heriot-Watt, we developed solutions for the automatic composition of web services, using AI planning, applied to gene expression databases. Our work on Argumentation Systems in the Life Sciences also originated as part of Sealife.
Further Details: [Sealife on BISEL pages]
[Sealife project web site - no longer online]